Project:Blood typing
How genes code for blood group
Blood group is determined by the combination of A and B antigens in your red blood cells. A and B individuals have only their respective corresponding antigens, AB individuals have both, and O individuals have none.
The production of these antigens is determined by the "histo-blood group ABO system transferase" gene (1062 base pairs), which is part of the ABO gene locus. This gene codes for the expression of a glycosyltransferase enzyme which by acting on another antigen(H), produces A or B antigens. The A and B allelic forms of the gene code for different forms of glycosyltransferase which affect the H antigen in different ways. O alleles code for another protein that doesn't affect the H antigen, meaning no A or B antigens are expressed. O alleles have a deletion of G at 258, while B alleles have a single nucleotide polymorphism (SNP) from G to A at position 700. The gene we're looking at is here: ABO gene. You need to set the origin to 29 under the tools menu, as this viewer is for the entire ABO gene; the histo-blood group ABO system transferase gene doesn't start until position 29.
Each person has two of these alleles, one from each parent. A and B are dominant, O is recessive, so the possible combinations are:
| Alleles | Blood group |
|---|---|
| AA | A |
| AO | A |
| BB | B |
| BO | B |
| AB | AB |
| OO | O |
In the UK the distribution of A, B, AB and O is 42%, 10%, 4% and 44%.
Process overview
1) Obtain two sequences of DNA through PCR, the first containing the deletion site at 258, and the second containing the SNP site at 700
2) Use restriction enzyme KpnI on the first fragment to cut the O alleles only, and use restriction enzyme AluI on the second fragment to cut the B alleles only. (KpnI's cutting site is GGTAC^C - this sequence is found in 0 alleles, but in A and B alleles the sequence is GGTGACC, hence they are not cut. AluI's cutting site is AG^CT - in A and O alleles the initial A is a G, hence only B alleles are cut.)
3) Do gel electrophoresis on the resulting fragments, hopefully resulting in distinguishable bands to show the 6 different alleles. We should then be able to determine an individual's blood type.
Process reality
Much of the necessary equipment we already have from the sex typing experiments. Of the new things, we need the restriction enzymes, and possible a new gel - see below.
All fragments in the papers are between 80 and 200 bp long. For this we would need a polyacrylamide gel (which we have decided against due to difficulty with handling) or a high quality agarose concentrated at 3%, which is a bit expensive, but not impossible. So we have decided to design our own primers to get larger fragments to work with.
Design of primers for larger fragments:
Using the ApE software, we have found a set of primers to enable us to use longer fragments. Here is the file with the primer sequences (Save this link as a file): ApE file, and Download ApE here
Primers for G deletion sequence:
| Primer | Sequence | Length | GC% | Tm(°C) |
|---|---|---|---|---|
| P1 forward | 17484 5' CCCGCAGGTCCAATGTTGAG 3' 17503 | 20 | 60 | 59 |
| P1 reverse | 18268 5' ATCTGACAGAGAAGTGACCACG 3' 18247 | 22 | 50 | 58 |
Product is 784bp.
After digestion with KpnI you get two sizes of fragments - 243bp and 541bp
Primers for G to A SNP:
| Primer | Sequence | Length | GC% | Tm(°C) |
|---|---|---|---|---|
| P2 forward | 19125 5' GAGGTGGATTACCTGGTGTGC 3' 19145 | 21 | 57 | 59 |
| P2 reverse | 19473 5' GCACCTTGGTGGGTTTGTGG 3' 19454 | 20 | 60 | 60 |
Product: 347 bp.
After digestion with AluI : two fragment sizes of 96 bp and 251 bp
New equipment
Sigma aldrich have KpnI and AluI. I couldn't find them on NBS bio. A cheaper alternative is to try to get them from NEB (http://www.neb.uk.com/).
If we do need high quality agarose here is sigma's selection. NBSbio have agarose but no info on its purity.
Sources
Most of the procedure came from this paper. With some more here. For other papers and background see here.
Here is a sequence viewer for the ABO gene. Histo-blood group ABO system transferase starts at position 28 - so to find the deletion at 258 and the SNP at 700 you have to add 28 to the numbers on the viewer. Go from the 5' end.
Extraction and PCR 15/8
- Extract DNA (two samples, Will and Nicholas) from cheek by rubbing with pipette tip and taking saliva
- Mix with 250ul of chelex 100, cover with a drop of oil and incubate at 56°C for 30 mins.
- Centrifuge for 5 mins and remove supernatant. Supernatant kept to view genomic DNA later.
- Two reactions prepared - Nicolas (AMELX/Y primers), Will (PB1 primers)
- Reaction volume - 12.5ul Taq, 5ul template, 5ul primers, 2.5ul deionised water.
- PCR: initial denaturation at 96°C for 5 mins, then 35 cycles of 96°C for 1 min, annealing at 55° for 30 secs, extension at 72°C for 1 min.
- Results. PCR failed. Will's DNA was the one using blood typing primers.
Extraction and electrophoresis of genomic DNA 15-17/8
- Add cheek scraping + saliva to 1/2 a PCR tube of chelex (half bead layer, half liquid layer). Mix
- Incubate at 56°C for 30 mins, vortex briefly, 94°C for 10 mins. (Should have centrifuged and) take supernatant.
- Add 5ul of loading buffer to 25ul of template - visualise on 1% agarose gel
- DNA visible in well.
Extraction and PCR 19-20/8/12 (Will)
- Chelex extraction. Visualised and saw glow in well so proceeded to use this as template.
- 25ul total reaction volumes. Quantities assumed as no record: 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC).
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From right:
- 1: 250-10000bp ladder (from hackspace). Ladder key
- 2: PC + amelogenin (sex typing) primers (combined aliquot)
- 3: PC + amelogenin (sex typing) primers (separate aliquots)
- 4: PC + PB1 primers
- 5: Template put through PCR program with no other reagents
- 6: Template + separate AMEL primers
- 7: Template + separate PB1 primers
- 8: Template - no PCR
Conclusions:
- As PC worked with PB1 primers but not AMEL, there is either a problem with the AMEL primers, or the annealing temperature of 55C doesn't work for them.
- Something in the reagents / template stops PCR working for our extractions OR there is too little DNA extracted. However even if the extracted DNA is very little, surely is enough for PCR? And we are pretty sure there is DNA there because it shows up genomically in the wells.
Extraction and PCR 03/10
- About 2ul of saliva scraped from cheek + 325ul chelex (half bead layer, half liquid layer). Mix.
- Incubate at 56°C for 30 mins (missed vortex), 94°C for 10 mins. Centrifuge and take supernatant.
- PCR mix: 5ul template, 5ul primers, 2.5ul dH20, 12.5ul Jumpstart taq readymix
- Positive control: 1ul template, 5ul primers, 6.5ul dH20, 12.5ul Jumpstart taq readymix
- PCR: 94C 2min, 30 cycles (94C 30s, 56C 30s, 72C 2min), 72C 5min.
From left:
- 1: 250-10000bp ladder (from hackspace). Ladder key
- 2: Positive control PCR product.
- 3: Mike PCR product (not there)
- 4: Simon PCR product (not there)
- 5: Will PCR product (not there)
- 6: MF1 (From UCL)
- 7: N.H / N.1+ (From UCL)
The positive control worked (You can see if you look closely). However the band was significantly weaker than the gel run on August 20th with same PC + same primers. None of the PCRs on our extractions worked.
I also ran our 3 genomic samples, and nothing was there - providing a reason for PCR failure.
Extraction 25/10 (Done by Will and Lui)
- 10ml dH20 vigorously rinsed in mouth (with cheek biting) for 3 mins
- Spat out, added ∼1.5ml washing up liquid, mixed, left for ∼10 mins.
- Split samples into two groups
1st group:
- Took ∼500ul sample, added ∼100ul Proteinase K solution
- Incubated for 1hr at 50C, 10 mins 94C
- Added 1 volume 100% EtOh, small amount of precipitate observed, so centrifuged at 13000 rpm for 20 mins
- Dried at ∼50C
- Added ∼350ul 70% IPA, centrifuged at 13000 rpm for 10 mins
- Air dried pellet
- Tried to resuspend in 500ul TE but pellet wouldn't resuspend completely. Unsure if none had dissolved or some had. Even incubating at 60C didn't work. Edit - seemed none dissolved as nothing showed up on gel. Didn't dissolve even in 10 ml dH20.
2nd group (remainder of sample - 11ml:
- Added 10 shakes meat tenderiser
- Added 1/2 volume 70% IPA. Significant amount of precipitate observed. Able to remove it with pipette.
- Dried at ∼50C
- Split, washed one with 70% IPA, centrifuged 10min at 13000 RPM. Then resuspended pellet in 500ul TE. Other only resuspended, no wash. Resuspensions didn't totally work (not sure if any dissolved).
PCR of 1st group 27/10 (Will's sample)
- 25ul total volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- PCR was Initial D - 95C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- Gel - loaded 5ul of ladder, 15ul of each sample, ∼ 0.2 vol loading buffer
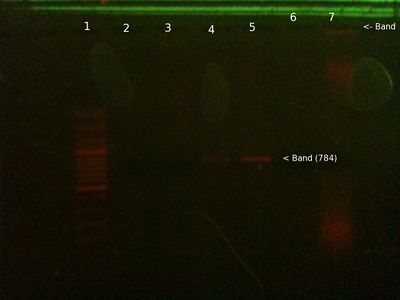
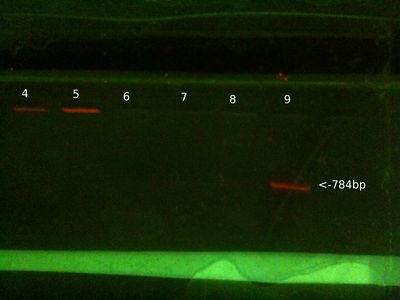
From right (after 35 mins at ∼ 80V:)
- 1: 250-10000bp ladder. Ladder key (At the far right, you can barely see it)
- 2: Will PCR product (PB1 primers) (Nothing)
- 3: Positive control PCR product (PB1 primers) - We expect this to be 784 bp, looks right.
- 4: Positive control PCR product (PB2 primers) - We expect this to be 347 bp, looks right.
- 5: Will genomic (Nothing)
Conclusions:
- Something went wrong with the ladder - it's barely visible. I used standard quantity so unsure why.
- My extractions failed PCR, and nothing showed up in genomic - hence extractions failed. If the pellet was DNA and had not degraded, it just must not have dissolved.
- Both controls worked - so both sets of our blood typing primers work.
Extraction 28/10
- Used this protocol. No DNA precipitated, so resolved to try again tomorrow using cheek swish method instead of cheek scrape.
Extraction 29/10
- Used this protocol. Instead of scraping cells from cheek, swished 5ml of dH20 in mouth for 3 mins while chewing cheeks. Then took 200ul of this and proceeded with extraction.
- One reaction had 1% Triton. 150ul of sample + lysis buffer added to 50ul PK. Incubated. No precipitate observed, but washed where pellet would be with 70% Etoh. Stored in freezer in 70% Etoh.
- One reaction had 10% Triton. 150ul of sample + lysis buffer added to 50ul PK. Incubated. No precipitate observed, but washed where pellet would be with 70% Etoh. Stored in freezer in 70% Etoh.
- Added 0.5ml dishwashing detergent to 'mouth swish remainder' (4.5ml). Added a few shakes meat tenderiser. Took ∼150ul, centrifuged and took supernatant. Added 0.25 vol PK and incubated. Precipitated and washed, small amount of DNA observed, tried to resuspend but wouldn't dissolve. Stored at -20C.
- Used IPA to precipitate the rest. Large pellet extracted, air dryed for 5 mins, then would not dissolve in 300ul TE at 65C. Stored at -20C.
Extraction and PCR 31/10 (Will)
- Extracted DNA from cheek using this protocol
- Ran PCR with PB2 primers on this sample + the third sample from the 29/10 + a positive control.
- 25ul total volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 95C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after ∼40 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key
- 2: Will PCR product (Chelex extraction) (Nothing there)
- 3: Will PCR product (Ethanol extraction) - We expect this to be 347 bp, looks right.
- 4: Positive control PCR product - We expect this to be 347 bp, looks right.
- 5: Chelex extraction genomic
- 6: Ethanol extraction genomic (Nothing there)
Conclusions:
This needs to be repeated to confirm, but it seems to show that the ethanol precipitated DNA worked in PCR, even though it looked like none had dissolved, while the chelex didn't. If at least one has worked that's good news. It's strange though that the chelex extraction shows up as genomic in the well, but failed PCR, whereas the ethanol extraction is the other way round. Possible that something in chelex mix inhibited PCR, while quantity of genomic was too little to show up for ethanol, but was enough for PCR. Possibility of mix up in pipetting.
Mailing list discussion here
PCR 1st Nov 2012 (Will)
Repeated PCR from 31 Oct to test conclusions.
- Same inputs except that the chelex solution was centrifuged and resulting pellet discarded before PCR. Also another sample from 29/10 used in addition to one from last time. PB2 primers used.
- 25ul intended total volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC). Actually put 6.5ul of dH20 in each sample by mistake, meaning total volume of 29ul for all except PC.
- Initial D - 95C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 40 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key
- 2: Will PCR product (Chelex extraction) - We expect this to be 347 bp.
- 3: Will PCR product (3rd Ethanol extraction from 29/10) - Nothing
- 4: Will PCR product (1st Ethanol extraction from 29/10) - Nothing
- 5: Positive control PCR product - We expect this to be 347 bp
- 6: Chelex extraction genomic
Lanes 7 and 8 not visible on photo
- 7: 3rd Ethanol extraction genomic (Nothing there)
- 8: 1st Ethanol extraction genomic (Nothing there)
Conclusions:
This could mean that I mixed up chelex and ethanol extractions for the run on the previous day, and chelex actually worked then too. Otherwise these results mean that chelex worked this time but not last time, either because I removed PCR inhibitors with the centrifuge, or because we avoided the problem of small reaction volume discussed previously. This would also mean that the ethanol precipitated sample has stopped working. Need to do more to confirm. Odd that chelex genomic showed such a strong band last time but none this time.
Mailing list discussion here
Extraction and PCR 04/11 (Will)
- Extracted DNA from cheek using this protocol. Pellet quantity was larger than previous time, as swished in mouth for 1 min. perhaps too large.
- Ran PCR with PB2 primers on new chelex extraction from 4/11 + old chelex extraction from 31/10 + third sample from the 29/10 + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 40 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key
- 2: Will PCR product (4/11 Chelex extraction) - Nothing there
- 3: Will PCR product (31/10 Chelex extraction) - Good band
- 4: Will PCR product (3rd Ethanol extraction from 29/10) - Weak band
- 5: Positive control PCR product - No band
- 6: Chelex extraction 4/11 genomic - band
Lanes 7 and 8 not visible on photo
- 7: Chelex extraction 31/10 (Nothing there)
- 8: 3rd Ethanol extraction 29/10 genomic (Nothing there)
Conclusion:
This time the new chelex extraction didn't work (but showed up as gDNA. Will want to try again, possibly used too much. 31/10 chelex worked as expected. Old ethanol worked too, which was a surprise. Positive control didn't work, seemed very small volume after PCR, possible reason.
Extraction and PCR 07/11 (Will, Nicholas, Adam, Jim)
- Nick, Adam and Jim extracted DNA from their cheeks using this protocol.
- Ran PCR with PB2 primers on these 3 + Will's chelex extraction from 4/11 + Will's chelex extraction from 31/10 + Will's third sample from the 29/10 + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 40 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key
- 2: Nick PCR product
- 3: Adam PCR product
- 4: Jim PCR product
- 5: PC PCR product - nothing (low volume after PCR. Perhaps evaporation)
- 6: Will PCR product (4/11 Chelex extraction) - nothing
- 7: Will PCR product (31/10 Chelex extraction) - nothing (low volume after PCR. Perhaps evaporation)
- 8: Will PCR product (29/10 3rd Ethanol precipitation)
- 9: Nick gDNA - small amount visible in well (but not in photo)
Lanes 10, 11 and 12 not in photo
- 10: Adam gDNA - visible in well
- 11: Jim gDNA - not visible in well
- 12: Will gDNA (4/11 chelex extraction) - small amount visible in well + smear running ahead of ladder (disappeared over time)
Conclusions:
- 2/3 of new chelex extractions worked, this is good. However they are very inconsistent despite using the same extraction protocol; Nick's is strong, Adam's faint, Jim's failed. Not sure why this is.
- The ethanol precipitation extraction from 29/10 continued to work. This means we can try to get more successes with ethanol.
- The two older chelex extractions didn't work. The 4/11 was expected to fail, as it failed last time. The 31/10 worked last time. Possible reason for failure this time was that volume was low coming out of PCR, with droplets on the side of the tube.
- Positive control failed again. This also had low reaction volume coming out of PCR like last time, and droplets on the side of the tube, meaning evaporation is a possible reason.
- Many samples had smears running ahead of them (these had gone by the time the photo was taken), which has not happened the last few times. This could be because this PCR was left overnight at 4C before the gel was run the next morning, leaving time for the DNA to degrade, whereas the last few gels were run straight away. Or (less likely) because unlike the last few times, autoclaved pipette tips were not used for the initial extractions.
- Some more in mailing list discussion
Extraction and PCR 11/11 (Will, Simon, Ben)
- Will, Simon and Ben extracted DNA from their cheeks using this protocol. Possibly too much DNA extracted (∼200ul pellet). Clumps remained once mixed with chelex (possibly also used too many beads for chelex + lots of mixing with pipette tip which may have broken up DNA), as could only get 150ul of supernatant at end. Supernatant seemed slightly cloudy. Quantity of Simon's extraction less than others.
- Ran PCR with PB2 primers on these 3 + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 5ul primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 35 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key
- 2: Simon PCR product (Faint)
- 3: Ben PCR product (strong)
- 4: Will PCR product (strong)
- 5: PC PCR product (Failed)
- 6: Simon gDNA (Very faint)
- 7: Ben gDNA (Strong)
- 8: Will gDNA (Medium)
Conclusions:
- All 3 extractions worked. Simon's was weaker than others. Possible reasons could be that his quantity of solid extracted from cheek wash was less, he had more contamination, or he did not mix the chelex solution as thoroughly.
- Positive control failed for the 3rd consecutive successful PCR. Implies it may have degraded.
- gDNA bands strength reflect Simon's less successful PCR.
PCR to test SYBR green Taq readymix 17/11
We did PCRs on DNA samples provided by Tom to test the SYBR green Taq. As you see below I made a mistake (I think) and did not add reagents as I should. However if everything works it should still tell us what we want.
- 25ul total reaction volume - 1ul template, 6.5ul dH20, 5ul primers, 12.5ul Taq readymix
- S1T1 - old PC old Taq
- S1T2 - old PC old Taq (should have been old PC new Taq)
- S2T1 - new PC new Taq (should have been new PC old Taq)
- S2T2 - new PC new Taq
After PCR stored at -20C.
Extraction and PCR (Will, Lui, Leo, Simon) 18/11
- Leo and Lui extracted DNA from their cheeks using this protocol.
- Will and Simon extracted DNA from 1 pea using the same protocol. Pea was crushed in a tube to make it a similar consistency as cheek extract
- Ran PCR with PB2 primers for human and pea primers for pea on these 3 + a positive control. Gel included PCRs from 17/11.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 35 mins at 80V:)
- 1: 250-10000bp ladder. Ladder key. Ladder wasn't visible. Not sure why
- 2: Leo PCR product (Nothing there - poss very faint band but not enough to be conclusive)
- 3: Lui PCR product (Nothing there)
- 4: PC PCR product (Weak band)
- 5: Pea PCR product (Diffuse band in roughly the right position - 280bp, but also roughly same position as pea gDNA)
- 6: S1 T1 (very weak band - pipetting error when loading sample into lane)
- 7: S1 T2 (weak band)
- 8: S2 T1 (weak band)
- 9: S2 T2 (weak band)
- 10: Pea gDNA (diffuse band at 0-300 bp)
Issues
- The agarose gel turned out to be partially solidified when we poured it into the tray, but we managed to reliquify it using a hotplate and a water pan.
- The tray is leaking and needs to be fixed.
- We have now run out of agarose.
Conclusions:
- Both our cheek extractions failed. Only reason I can think of is that in previous experiments small quantities of solid from cheek was less likely to work than large quantities. However too large a quantity has also failed.
- Pea PCR product was roughly in the right place. However as band was diffuse and in the same place as pea gDNA I suspect it was in fact pea gDNA. Need to try again, perhaps with less template, perhaps with different PCR program.
- All PC test samples worked, indicating we can use the SYBR green taq.
Extraction and PCR (Will, Simon) 8/12
- Will and Simon extracted DNA from their cheeks using this protocol.
- Ran PCR with PB2 primers on these 2 + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul SYBR green Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 35 mins at 80V:)
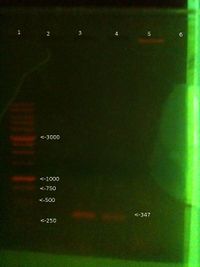
- 1: 100-3000bp ladder. Ladder key.
- 2: Will PCR product (Faint band at right position - 347 bp)
- 3: Simon PCR product (No band, or possible very faint band)
- 4: PC PCR product (Good band at right position - 347bp)
- 5: Will gDNA (Good band)
- 6: Simon gDNA (Good band)
Conclusions:
- Pipetting error loading Simon's sample could explain lack of band. Simon's quantity of pellet from cheek was larger than Will's.
- Will's PCR worked, but faintly. We need to focus on improving PCR consistency and resolution.
PCR (Will, Lui, Tom) 12/12
- Redid PCR on Will's extraction from 8/12 to see if doubling quantity of template helped. PCR 1 was same as last time, PCR 2 was with twice the template and no dH20. First PC was same as last time, second was with twice the template and corresponding less water.
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
From left (after 30 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Will PCR 1 product (Faint band)
- 3: Will PCR 2 product (No band - although band was there earlier in PCR)
- 4: PC PCR 1 product (Good band)
- 5: PC PCR 2 product (Good band)
- 6: Will gDNA (Faint band)
Conclusions:
- Doubling quantity of template doesn't help, actually made band weaker. This indicates either that too much template hurts the reaction, or that template contains inhibitors - so doubling template doubles inhibitors.
- gDNA band was weaker than last time. Probably degradation.
Extraction and PCR (Will, Jim, Mike) 10/1/13
- Will Jim and Mike extracted DNA from their cheeks using this protocol. (15 secs swishing)
- Jim and Mike's pellet was much smaller than Will's
- Ran PCR with PB2 primers on these 3 + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul SYBR green Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- PCR left overnight. 15ul of samples loaded, 6ul of ladder loaded.
From left (after 35 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Jim PCR product (No band)
- 3: Mike PCR product (No band. Blob is result of accidentally touching gel with dirty pipette)
- 4: Will PCR product (No band)
- 5: PC PCR (band)
- 6: Jim gDNA (No band)
- 7: Mike gDNA (No band)
- 8: Will gDNA (Faint band)
Conclusions:
- No or not enough DNA extracted. Possibly fault of 'whirly' technique. Could also be due to leaving overnight.
- Poor resolution. Possibly due to new TBE.
Extraction and PCR (Will, Simon) 13/1/13
- Will and Simon extracted DNA from their cheeks using this protocol. (30 secs swishing and cheek chewing). Centrifuge for 5 minutes. Solution with chelex stirred every few minutes during 100C incubation.
- Ran PCR with PB2 primers on these two + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul SYBR green Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- 15ul of samples loaded, 6ul of ladder loaded.
From left (after 40 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Simon PCR product (nothing)
- 3: Will PCR product (faint gDNA band, not visible in photo)
- 4: PC PCR product (nothing)
- 5: Simon gDNA (band, not visible in photo)
- 6: Will gDNA (band)
Conclusions:
- Seems to have been a PCR failure rather than extraction failure. No idea why, but most likely problems with taq or primers, or pipetting error. Also the template had a yellowish tinge, which is something past PCR failures have also had.
PCR (Will, Simon) 14/1/13
- Repeat PCR on samples from 13/1. Samples centrifuged to remove debris, but yellowish tint remained.
- Ran PCR with PB1 primers on these two + 1 positive control with SYBR green Taq readymix + 1 PC with Taq readymix
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul SYBR green Taq readymix / Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- PCR left overnight. 15ul of samples loaded, 10ul of ladder loaded.
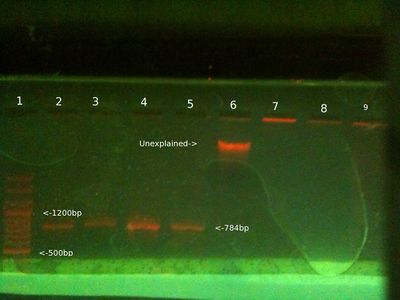
From left (after 35 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Simon PCR product (Nothing)
- 3: Will PCR product (Nothing)
- 4: PC SYBR green Taq PCR product (Faint band)
- 5 PC Taq PCR product (Band)
- 6: Simon gDNA (Nothing)
- 7: Will gDNA (Faint band in well + unexplained blobs)
Conclusions:
- This time PCR worked on the PCs (better on straight Taq readymix than SYBR green Taq readymix). So something wrong with extractions. Either contamination or degradation (gDNA bands weak or non existent compared to same samples 24 hours earlier)
PCR (Nicholas, Ben) 16/1/13
- Repeated PCR on successful samples from November 2012.
- Ran PCR with PB1 primers on these two + 1 positive control with SYBR green Taq readymix
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul SYBR green Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- PCR left overnight. 7ul of samples loaded, 7ul of ladder loaded. (Less than previous times)
- Ran electropheresis with new, larger power source, with constant 80V.
- After 5 mins: bands were initially visible, especially ladder, Ben, and PC.
- After 40 mins: We left it for too long, and the bands had disappeared. No photo taken.
Notes: We could work on a better visualisation system. Currently using black pouch as background. The gel bath has broken, we will try a plastic food container next time. The main button on the smaller pipette has broken, super glue needed.
Extraction and PCR (Will, Simon, Lui) 12/2/13
- Lui extracted DNA from his cheek using this protocol.
- Ran PCR with PB2 primers on this + 3 RCA extractions + a positive control.
- 25ul total PCR reaction volume - 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul Taq readymix, 2.5ul dH20 (6.5ul for PC)
- Initial D - 96C 5 mins, then 35 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- 15ul of samples loaded, 3ul loading buffer, 7ul of ladder loaded.
From left (after 40 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Lui PCR product (nothing)
- 3: Pip PCR product (nothing)
- 4: A PCR product (band)
- 5: J PCR product (very faint band)
- 6: Positive Control PCR product (band)
- 7: Lui gDNA (nothing)
- 8: Pip gDNA (strong band)
- 9: A gDNA (faint band)
- 10: J gDNA (band)
Conclusions:
- Lui's extraction didn't contain enough liquid after boiling step, so he added more water afterwards and mixed. This could be responsible for extraction failure. Not sure reasons for variations in RCA extractions.
Extraction and PCR (David, Victor, Will) 17/2/13
- David and Will extracted DNA from cheeks using this protocol.
- Ran PCR with PB1 primers on this + Victor's extraction from 13/2 (same protocol) + a positive control.
- 25ul total PCR reaction volume - 2.5ul dH20 (6.5ul for PC), 5ul template (1ul for PC), 2.5ul each forward & reverse primers, 12.5ul Taq readymix
- Initial D - 96C 5 mins, then 40 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- 25ul of PCR samples loaded, 15ul of gDNA samples loaded (2ul for PC gDNA). 2ul loading buffer, 5ul of ladder loaded.
- We expect to see PCR bands at 784bp.
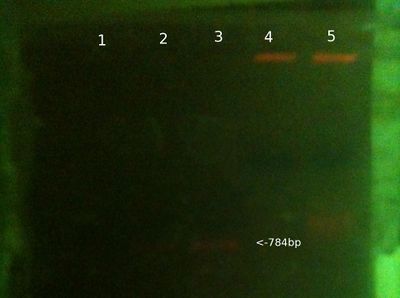
From left (after 30 mins at 80V:)
- 1: 100-3000bp ladder. Ladder key.
- 2: Victor PCR product
- 3: David PCR product
- 4: Will PCR product
- 5: PC PCR product
- 6: PC gDNA
- 7: Victor gDNA
- 8: David gDNA
- 9: Will gDNA
Notes/Conclusions:
- Increased PCR cycles from 35 to 40, and increased amount of product loaded onto gel. These could explain results being better than normal.
- Odd that PC gDNA travelled further in gel than our extractions.
- Also more confirmation that there's seemingly no correlation between strength of gDNA and strength of PCR product.
Extraction and PCR (JJ, Bene, Will) 20-23/2/13
- JJ and Bene extracted DNA from cheeks on wed 20th using this protocol. Small amount of cheek cells for each.
- On 23rd ran PCR with PB1 primers on these + a positive control.
- 50ul total PCR reaction volume - 5ul dH20 (13ul for PC), 10ul template (2ul for PC), 5ul each forward & reverse primers, 25ul Taq readymix
- Initial D - 96C 5 mins, then 40 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- 20ul of PCR samples loaded, 20ul of gDNA samples loaded. 4ul loading buffer.
- 1: JJ PCR product (no band)
- 2: Bene PCR product (very faint band)
- 3: PC PRC product (band)
- 4: JJ gDNA (band)
- 5: Bene gDNA (band)
Notes/Conclusions:
- PCRs either failed or were too weak to show. Differences in this reaction that could have affected the result were: we doubled PCR quantities, then loaded 20ul from the result. gDNA bands were fine, so strange PCR didn't work as PC was ok (but not strong).
Extraction and PCR repeat
- Bene did a new extraction using same protocol.
- Ran PCR with PB1 primers on this + JJ's old sample + Will's sample from 17/2 (1 with 50ul total volume, 1 with 25ul).
- 50ul total PCR reaction volume - 5ul dH20 (13ul for PC), 10ul template (2ul for PC), 5ul each forward & reverse primers, 25ul Taq readymix. (Halve quantities for Will2)
- Initial D - 96C 5 mins, then 40 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- 20ul of PCR samples loaded, 3ul loading buffer. Same gel used hence lanes 4+5 same as that one.
- 4: JJ gDNA (band)
- 5: Bene gDNA (band)
- 6: JJ PCR
- 7: Bene PCR
- 8: Will PCR (50ul)
- 9: Will PCR (25ul)
Notes/Conclusions:
- This seems to indicate pretty clearly that doubling the reaction quantity caused the PCRs to fail. Will try again on JJ's and Bene's with 25ul.
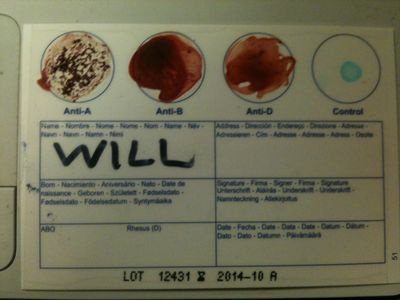
Blood test
We bought 5 home testing kits to check our results. Will is A-, as shown here. Each circle contains different types of antigens, and if your blood agglutinates in a circle it means you have that antigen.
PCR and Restriction digest (Simon, Lui, Will) 26/2/13
- Ran PCR with PB1 primers on JJ's sample from 20/2 and Bene's sample from 23/2.
- Did two reactions for each gDNA sample. 25ul total PCR reaction volume - 5ul dH20, 10ul template, 2.5ul each forward & reverse primers, 12.5ul Taq readymix
- Initial D - 96C 5 mins, then 40 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- Ran samples on gel to check PCR succeeded. JJ's did, couldn't tell with Bene's as we fucked up loading it onto the gel.
Restriction digest
- Incubated JJ's PCR product with Kpn1 at 37C for 65 mins. Restriction digest mix contained 25ul PCR product, 1ul Kpn1, 15ul buffer.
- Ran result on a gel. PCR product remained undigested.
Notes/Conclusions:
- We'll need to vary the quantities of the restriction digest to try and get better results.
PCR and Restriction digest (Tom, Will) 27/2/13
- Ran PCR with PB1 primers on JJ's sample from 20/2 and Bene's sample from 23/2.
- Did two reactions for each gDNA sample. 25ul total PCR reaction volume - 5ul dH20, 10ul template, 2.5ul each forward & reverse primers, 12.5ul Taq readymix
- Initial D - 96C 5 mins, then 40 cycles of D-96C 1 min, A-55C 30 secs, E-72C 1 min.
- Incubated one of each PCR product with Kpn1 at 37C for 90 mins. Restriction digest mix contained 25ul PCR product, 1ul Kpn1, 3ul buffer, 3ul BSA.
- Bene's samples: neither PCR product nor digest product showed, indicating PCR failed.
- JJ's samples: PCR product showed up, digest product didn't, indicating either digested product was too little to see, or something destroyed the DNA during digestion, or there was inconsistency between JJ's two PCR products
Notes/Conclusions:
- If we assume JJ's digested product was too little to see, we need to work on a better imaging system, or get more concentrated DNA out of PCR.